Protein-protein interactions play important roles in both inter- and intra-cellular signaling [1]. Among the various methods of detecting protein-protein interactions, co-IP is cost-saving, easy to handle, and can be applied to various types of sample, such as plant cells [4], mammalian cells [5] yeast [2], and nematode tissue [3].
The figure below shows the workflow of co-IP.
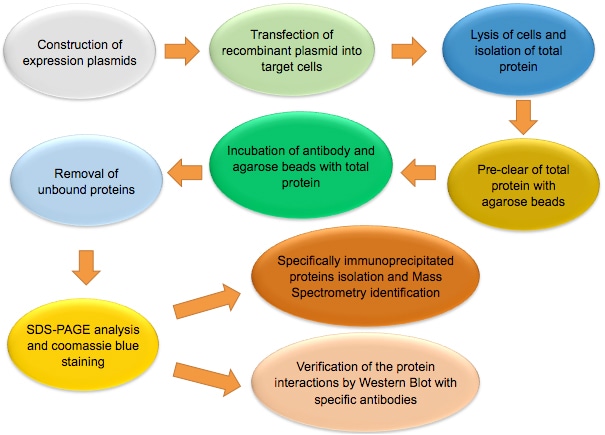
Note: Co-IP is a multi-step assay. Experiment should be performed under non-denaturing conditions. Not only high quality reagents, but also careful planning and rigorous operation are needed to obtain good result. Each step of the experiment, from sample preparation, antibody-beads incubation, to co-IP elution, should be tightly controlled to ensure success of the experiment.
Please refer to the relating protocols on our website for the following expression plasmid transformation, colony screening and positive colony verification steps.
Cell type used for transfection is determined according to the requirement of the experiment.
Note: In order to maintain the physiological state of the proteins, manipulation of the cell lysis should be under low temperature, the cell lysis buffer should contain mild non-ionic detergent, proteinase inhibitor, and glycerol.
Choice of detergent is essential for the success of the co-IP assay. Different types and concentration of detergents can affect co-IP result in the following aspects:
Alternatively, the antibody can be added for a few hours prior to the addition of Protein A/G agarose beads.
Note: In general, 1 μg antibody is used for 1 mg total protein. The usage of antibody should be no more than 5 μg, since too much antibody addition may generate false positive result.
Proper control is important. In general, incubation with an irrelevant protein antibody is usually used as negative control. Besides, incubation of antibody with bait protein free total protein solution can also serve as negative control, especially when polyclonal antibody is used.
Tagged target proteins can also be applied to co-IP assay, and corresponding anti-tag antibodies are needed.
Illustration of co-IP of Arabidopsis Thaliana Membrane Proteins [4]
![Illustration of co-IP of Arabidopsis thaliana membrane proteins [4]](/wp-content/themes/profacgen/img/Co-IP-4.jpg)
Q1: Western Blot result indicates low expression levels of target proteins.
S1: Increase the amounts of reagents, such as plasmids.
Q2: Coomassie blue stain shows weak staining of target proteins.
S2: This is likely due to the loss of proteins during the co-IP process. Optimize the co-IP conditions by using less stringent cell lysis buffer, more antibody-conjugated beads, or centrifuge at higher speed and/or for longer time during washing.
Q3: Strong target protein bands, but few prey protein bands.
S3: Restart with more cells to collect larger volume of cell lysate, improve the co-IP conditions, and pay special attention to buffer pH.
Q4; Strong target protein staining and clear prey protein bands, but high background in controls.
S4: Follow rigorous co-IP washing process, pre-clear total protein by adding greater amount of control agarose beads, reduce the concentration of detergent.
Q5: High noises of MS analysis result.
S5: Follow rigorous gel operation process, excise bands with margins as small as possible.
Q6: Over-abundant keratin signals.
S6: Handle gels with extreme caution to avoid direct skin contact with gels.
References:
Fill out this form and one of our experts will respond to you within one business day.