Membrane proteins constitute one third of all genes in the human genome, and they play critical roles in cells as receptors, enzymes, ion channels, and transporters. Owing to their functional importance, membrane proteins account for more than 50% of current drug targets. However, experimental structural data are sparse, mostly due to difficulties in their overexpression, reconstitution into membrane mimetics, and subsequent structure determination. The lack of high-resolution structures hinders drug design process and the engineering of receptors with novel function for biotechnological applications, resulting in high expectations for computational modelling tools to help fill this gap.
Computational structural modeling represents an important alternative approach to obtaining three-dimensional membrane protein models at the atomic level. With the growing amount of available high resolution membrane protein structures, the template situation for homology modelling procedures is improving quickly, which ensures high accuracy of resultant models. The workflow of predicting the structure of membrane proteins is basically the same as for their soluble counterparts, which starts with sequence alignment of the target protein with the selected template(s). During the model building process, the membrane-spanning regions are predicted as the core, whose structure is built and refined first. This is followed by the prediction of loop conformations, especially for the extra-membranous regions, to generate a complete model.
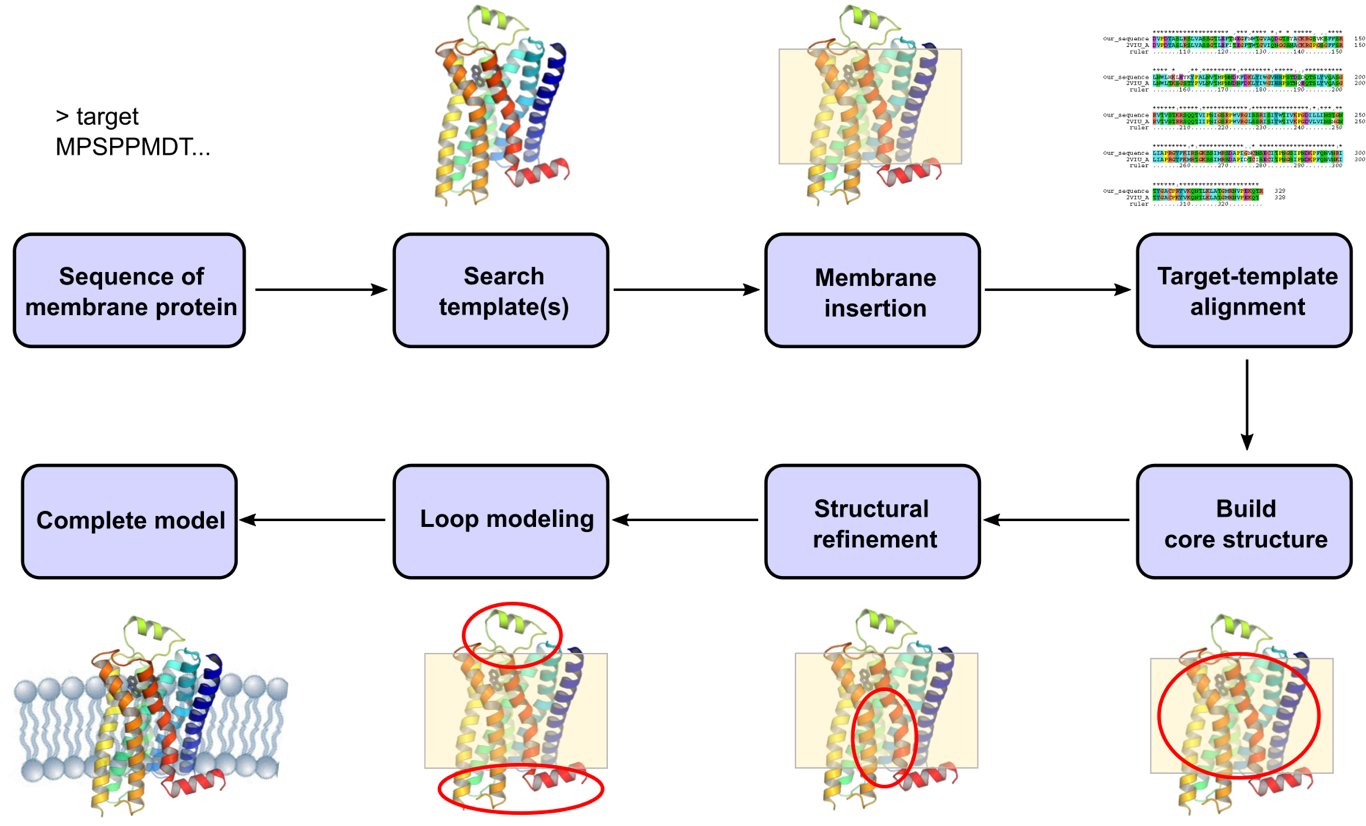
Profacgen adapts conventional homology modeling methods with several membrane-protein-specific steps to achieve better modeling results by avoiding any bias towards soluble globular proteins in the algorithm. In cases where structural information from closely related proteins is unavailable, distant structural homologues are exploited for the reconstruction of the core and loop regions. Over the years, we have accumulated extensive experiences with the modeling of various types of membrane proteins, including G protein-coupled receptors (GPCRs) and transport proteins. The resultant structural models are all quality verified and can be used for computational drug design, molecular dynamics simulation, as well as directing further experimental work such as structure-based protein engineering.
Our goal is to predict the structure of membrane proteins from their amino acid sequences with an accuracy comparable to experimental approaches, overcoming many difficulties in the structure determination of membrane proteins. We customize the service according to the specific requirements from the customers. Please do not hesitate to contact us for more details about our membrane protein modeling service.
Fill out this form and one of our experts will respond to you within one business day.