Recombinant proteins are widely used in biological and biomedical research[1], and recombinant protein expression has become a commonly used tool. Nevertheless, there are still many aspects to consider before starting an expression project. For example, which expression system is chosen for obtaining the protein of interest? Should it be expressed in bacteria, in yeast, in insect cells or in mammalian cells? What is the best expression vector to be used? As to the bacterial expression system, which strain should be chosen? Is there any differences in the expression condition between the full-length protein and a protein fragment? Which affinity tag should be used to facilitate both protein expression and subsequent purification steps? How to work out an appropriate purification strategy? Since protein property can vary a lot, all conditions need to be carefully chosen to fit the project requirements.
Profacgen is equipped with experienced scientific team and well-developed recombinant protein expression systems to serve the biopharmacetutical and life science field fully customized recombinant protein production services. Our team can help customers work out a best fit expression plan and purification strategy for the target protein with its intended application. Over the past 10 years, our Profacgen team has targeted and purified thousands of proteins from the prokaryotic systems and eukaryotic system with both structural integrity and functional activity.
Of all the expression hosts, E.coli is usually the first choice when applicable because of its ease of manipulation and low cost.

Profacgen has extensive experience in optimizing E.coli host strain and expression vectors for more efficient recombinant protein production. Various proteins, from prokaryotic proteins to eukaryotic proteins, from enzymes to transcription factors, from full-length to truncated domains, from monomeric protein to large protein complexes can be produced in our specialized E.coli expression system. For those proteins that do not express in soluble form due to misfolding in the E.coli system, we also have developed strategies to extract them from the inclusion bodies.
For high-level protein production, BL21(DE3) is a basic and most widely used E.coli strain. It has advantages of being deficient in both lon and ompT proteases and is compatible with the T7 lacO promoter system [2]. Our scientists have also developed several BL21 derivatives carrying additional helper plasmid to overcome the bad effects of codon bias in protein expression.
1. Small-scale expression evaluation & expression optimization
Our service allows you to evaluate your target protein expression in your chosen expression system; identify the construct and conditions that gives you the most robust soluble expression of your target protein. We can work with you to design the expression trial according to your exact requirements. Validation of soluble protein expression in a chosen E.coli strain generally takes 1-2 weeks.
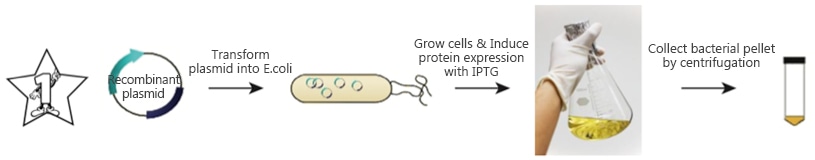
2. Standard Procedure:
a. Molecular cloning:
The gene of interest is inserted into a chosen expression vector. Usually the target protein is fused to a protein tag to facilitate expression and purification (i.e., 6*Histidine). The protein tag can be inserted to the N-terminus or C-terminus of the target gene for fusion protein expression [3]. Choice and position of the protein tag usually depends on the property of the target protein, expression condition and the customer’s application. A protease cleavage site can be inserted between the target protein and the protein tag, which will allow removal of the protein tag after purification. In addition, the expression vector also contains antibiotic resistance markers for clone selection [4].
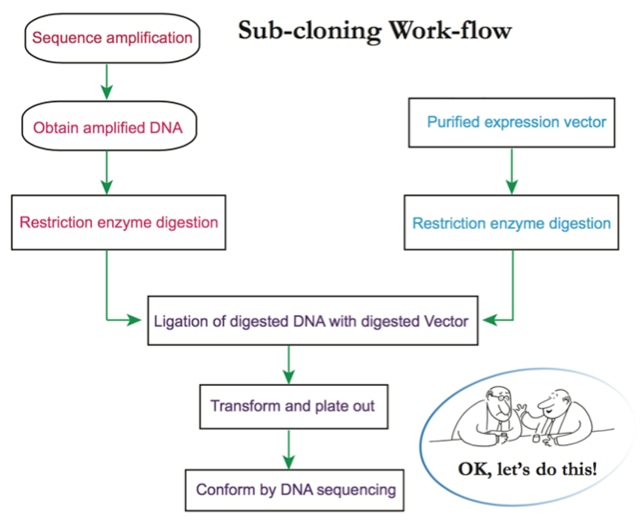
b. Transformation of the expression vector into chemically competent cells.
Competent cells are cells that can readily take up foreign DNA that, in this case, contain the target protein for expression. Below is a standard protocol for most expression vectors and cell lines [5].
| i. | Add the target vector into thawed competent cells. |
| ii. | Incubate the cells on ice, then treat with heat-shock and ice-shock. |
| iii. | Add liquid medium into treated competent cells for recovery. |
| iv. | Plate the transformed cells onto agar plates containing corresponding antibiotic for clone selection. |
c. Expression tests.
In order to identify the optimal conditions for growth and expression of the target protein, we perform complicated expression trials of temperature, induction time and expression time.
| i. | Transform the chosen E.coli strain competent cells for expression with the constructed expression vector. |
| ii. | Select the appropriate colony by antibiotic screening. |
| iii. | Incubate the correct colony harboring the recombinant plasmid in LB until OD600 reached to a certain value. |
| iv. | Divide the culture, equilibrate at different chosen temperatures, and induce expression by addition of isopropyl-β-D-thiogalactoside (IPTG). |
| v. | Continue cell growth and take samples at different time points after induction. |
| vi. | The remaining cultures were allowed to continue growing and eventually harvested. |
| vii. | All the samples were then lysed by sonication and then centrifuged to separate the supernatant and pellet for sodium-dodecyl sulphate polyacrylamide gel electrophoresis (SDS-PAGE) analysis. |
| viii. | Subject the re-suspended cells to a single freeze-thaw cycle prior to DNase digestion. |
| ix. | Centrifuge the lysed cell to remove the cell debris and filter the supernatant through a filter membrane. |
Please note that parameters such as OD600 value prior to IPTG induction, IPTG concentration, expression temperature and harvest time point can be adjusted to the expression status of the recombinant protein.
d. Protein purification
Preparation of the bacterial lysate can be critical for the subsequentprotein purification. The right cell lysis condition can maximize recombinant protein extraction, minimize unwanted proteolysis and sample contamination with genomic DNA and protein oxidation [1].
| i. | Cell lysis. Add lysozyme to the lysis buffer during mechanical lysis by sonication or lysis by freeze-thaw procedure. PH and salt concentration of the lysis buffer needs to be optimized to enhance protein solubility and stability according to the physiochemical property (solubility and pI, etc.) of the target protein. Protease inhibitors and reducing agent such as Tris (2-carboxyethyl) phosphine hydrochloride (TCEP) are used to protect target protein from complicated cell lysate mixture. |
| ii. | Protein purification by chromatography. One huge advantage of recombinant protein is that purification is greatly simplified by the addition of protein purification tags. Specific binding of certain metal or antibody of the protein tags can therefore be used in affinity chromatography to achieve strong and specific binding of the target protein.
|
Click here to contact us for more technical information.
[1]Structural Genomics C, China Structural Genomics C, Northeast Structural Genomics C, et al. Protein production and purification[J]. Nat Methods, 2008, 5(2): 135-46.
[2]Studier F W, Rosenberg A H, Dunn J J, et al. Use of T7 RNA polymerase to direct expression of cloned genes[J]. Methods Enzymol, 1990, 185: 60-89.
[3](2) M-Q D P a R H G B. Mutation of the Catalytic Cysteine in Anopheles gambiae Transglutaminase 3 (AgTG3) Abolishes Plugin Crosslinking Activity without Disrupting Protein Folding Properties[J]. Journal of Emerging Investigators, 2014.
[4]Davies H A. Expression and characterisation of cardiovascular amyloid proteins.[D]. University of Liverpool, 2013: 288.
[5]Sambrook J, Fritsch, E. F., Maniatis, T. Molecular Cloning - A laboratory handbook.[M]. In: NOLAN, C. (ed.) 2 ed.: Cold Spring Harbour Laboratory Press., 1989.
Fill out this form and one of our experts will respond to you within one business day.