For structure-based virtual screening (SBVS), candidate molecules are computationally docked into 3D structure of the biological target that are derived from biophysical methods (e.g. X-ray crystallography, NMR spectroscopy, and cryo-electron microscopy), homology modeling, or molecular dynamics simulations. These docked compounds are then ranked based on their predicted binding affinity or complementarity to the binding site, as well as other criteria. Usually only a few top-ranked compounds are selected as candidates for further experimental assays. Profacgen SBVS services cover all the procedures ranging from initial stages of the process that include receptor and library pre-processing, to docking, scoring, and postprocessing of top scoring hits.
At Profacgen, our expertise applies molecular dynamics in SBVS to explore the likely poses of a compound, to rescore molecules, to investigate the importance of water molecules, the pathways of interactions (e.g. kinetic of binding, unbinding event), and to investigate target flexibility, explore binding pockets and/or discover cryptic pockets and to rationalize allosteric events. We also use different types of postprocessing steps such as consensus docking and scoring, rescoring with more rigorous binding free energy calculations or with machine learning scoring functions. SBVS has become an essential tool in assisting fast and cost-efficient drug discovery and lead optimization.
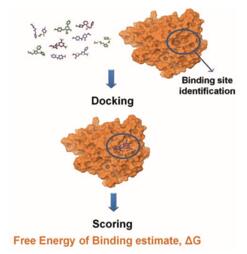
Figure 1. Docking and scoring in structure-based virtual screening. (Curr Top Med Chem. 2014)
Our company provides highly efficient SBVS services by fast and accurate molecular docking and scoring procedures. The compounds or fragments from a databased are docked into a binding site (or over the entire surface or in several binding sites or taking into account distance constraints such as covalent docking) and ranked by scoring function (e.g. force field based, empirical or knowledge based). Other SBVS services including fragment-based de novo ligand design and receptor-based pharmacophore screening are also available at Profacgen.
Our advantages
[1] Singh, N.; Chaput, L.; Villoutreix, B. O., Virtual screening web servers: designing chemical probes and drug candidates in the cyberspace. Brief Bioinform 2020.
[2] Lionta, E.; Spyrou, G.; Vassilatis, D. K.; Cournia, Z., Structure-based virtual screening for drug discovery: principles, applications and recent advances. Curr Top Med Chem 2014, 14 (16), 1923-38.
Fill out this form and one of our experts will respond to you within one business day.