Profacgen provides professional service for protein pull down assay. We have technical service group and most advanced instruments to perform the entire procedure with high-efficiency and high-quality.
Background
Pull down assay is an in vitro method to detect protein-protein interaction. The commonly used bait protein is a purified GST-tag protein. Pull down assay is usually followed by SDS-PAGE and Mass Spectrometry (MS) analysis to identify the interactor, and further genetic approaches or Western Blot analysis can be implemented to confirm the direct interaction.
Feature
The following figure shows the feature of pull down assay.
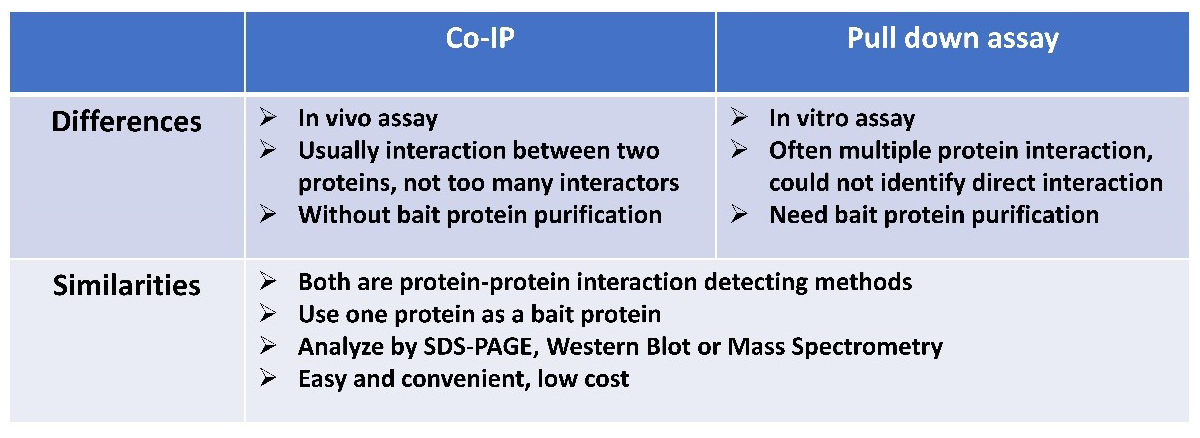
Comparison Between Pull Down Assay and Co-IP
Experimental design
Sample
You can provide us crude tissue sample or cell sample.
Tag preference
GST tag is the most preferred for bait protein purification, you can also use biotin tag or maltose binding protein (MBP) tag.
His-tag is not available, since the metal affinity beads will bind proteins non-specifically.
Control
All necessary control groups will be included according to the following table to verify interaction specificity.
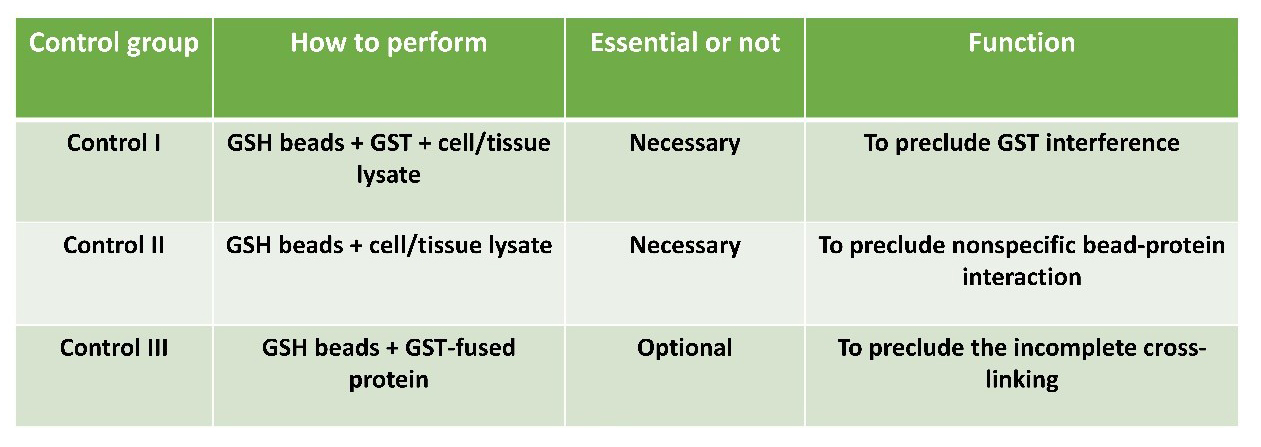
Customized service:
Experimental design
We will elaborately design our experimental scheme according to the protein of your interest, for example, its structure and function domain, cellular function, enzymatic or other activity, etc.
Specificity
Sometimes control samples are needed to increase the specificity of pull down assay. You can provide us a series of truncations, deletions, mutations of your purified tag fusion protein, or even related proteins from other genes.
Whether cross-link is needed
We will include the cross-linking (tagged protein with beads) step into the entire procedure or not, based on your requirement. The additional cross-linking step will make the final analysis more easily: since the tag fusion protein is covalently linked to the bead and will not appear in the subsequent gels. But please note: loss of some or all biological activity of specific proteins (particularly enzymes) may occur as a result of the chemical modification that occurs during cross-linking.
Optical scheme for sample treatment
For different purposes of experiments, we have different treatment methods for your samples (cytosolic protein extraction, peripheral protein extraction or membrane protein extraction).
Flowchart of the Pull Down Assay:
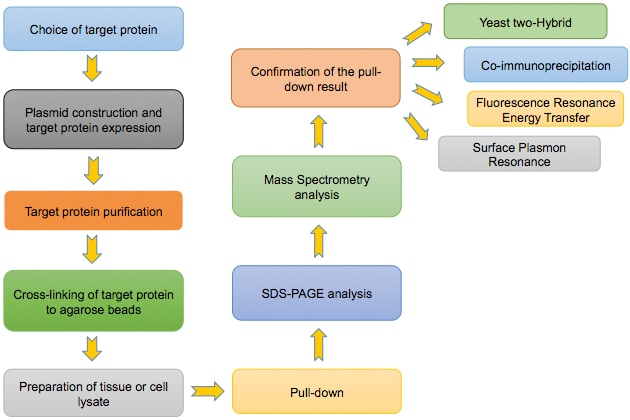
Construction of GST-tagged protein expression plasmid
| i. | Choose proper expression vector according to the target protein information provided by customers. |
| ii. | Insert the DNA sequence of target protein into the vector to generate recombinant expression plasmid. |
| iii. | Sequence the recombinant plasmid to assure no mutation in the sequence. |
| iv. | Transform the recombinant plasmid into E. coli component cell, and screen for positive clones. |
GST-fused protein expression and purification [1]
| i. | After positive colony verification, extract the plasmid and transform into E. coli BL21 for target protein expression. |
| ii. | Pick single colony into LB media with proper antibiotics, culture overnight. |
| iii. | Enlarge cultivation, to reach early exponential phase, add inducer to induce GST-fused protein expression (inducer free group as control). |
| iv. | Harvest bacterial cells, and resuspend with ice-cold cell lysis buffer. |
| v. | Pass the bacterial suspension through two freeze/thaw cycles. |
| vi. | Sonicate on ice to rupture bacterial cells and break up any DNA. |
| vii. | Add appropriate amount of DNase I and Triton X-100 into the cell lysate, incubate for 30 min at 4 °C with rotation. |
| viii. | Centrifuge the sample for 30 min at 4 °C. At the same time, treat the glutathione (GSH)-agarose beads with a small quantity of lysis buffer, centrifuge and carefully remove the supernatant. |
| ix. | Add the supernatant of lysate into pre-treated beads, incubate for 1 h at 4 °C with rotation. |
| x. | Centrifuge the mixture for 5 min at 4 °C, carefully decant the supernatant from the beads. |
| xi. | Rinse the beads with wash buffer, centrifuge for 5 min at 4 °C. Repeat for 3-4 times. |
| xii. | Add proper volume of bead storage buffer to the beads and store at -20 °C. The beads should never be allowed to freeze, or else the beads will crack. 50% glycerol can be added in the bead storage buffer to avoid this. |
| xiii. | Perform SDS-PAGE and stain with coomassie blue to analyze purification and quantitate the amount of protein bound to GSH beads (including inducer (+), inducer (-), supernatant, pellet, flow through, and beads sample). |
Normalization of proteins bound to the beads is necessary when different GST fused proteins are used as bait proteins.
Cross-linking of the target protein to beads
| i. | Add required volume of GST fusion protein beads to be cross-linked into proper size plastic column. Wash the column with PBS buffer twice. |
| ii. | Cross-link the GST fusion protein beads and column with cross-linker, incubate for 45 min at room temperature with continuous mixing. |
| iii. | Drain the column. |
| iv. | Add appropriate volume of quenching buffer to the column, gently mix for 10 min at room temperature (Block amine-reactive sites). |
| v. | Thoroughly wash the beads at 4 °C, store for use. |
Prepare GST fusion proteins and tissue lysate
| i. | Prepare recombinant GST fusion proteins attached to GSH-agarose beads and store at -20 °C in bead storage buffer for up to 6 months. |
| ii. | Prepare a tissue lysate (choose lysis method suitable to different protein locations: cytosolic, peripheral, or membrane). |
Ensure that lysate is free from even small traces of particle contamination. Contamination will result in large amounts of nonspecific protein on the gel. If present, it is very important to remove them by high speed centrifugation.
Pre-clear lysate with GSH beads
| i. | Wash a batch of (according to your sample amount) fresh or recycled GSH-agarose beads with wash buffer, centrifuge for 5 min at 4 °C, remove the supernatant and collect the beads in a minimal volume of buffer. |
| ii. | Combine the washed GSH beads with tissue lysate and incubate for 30 min at 4 °C. (Pre-clear the tissue lysate with the washed GSH beads for removal of endogenous GSTs present in the lysate and nonspecifically binding proteins) This will remove most of the endogenous cellular GST. |
| iii. | Collect the beads by centrifuge, and carefully pour off the supernatant into fresh, cooled tubes on ice. (Pre-cleared tissue lysate) |
| iv. | Repeat. |
This is especially important for tissue lysate that are rich in endogenous GSTs (particularly cytosolic extracts and lysates of specific tissues, for example, testis or liver).
Bind tissue lysate to the GST fusion protein beads
Add appropriate volume of pre-cleared lysate to corresponding volume of GST fused protein beads. Incubate for 1 h at 4 °C with rotation.
Recover and wash beads
| i. | Centrifuge the lysate and beads mixture at 4 °C, carefully remove the supernatant. |
| ii. | Wash the beads with ice-cold wash buffer, centrifuge and discard the supernatant. |
| iii. | Repeat for 5-6 times. |
Elute proteins for SDS-PAGE analysis
| i. | Add 1×SDS loading buffer into the beads, and place in an 85 °C water bath for 5 min. |
| ii. | Centrifuge the sample, store on ice for immediate use or in the freezer for future analysis. |
| iii. | Analyze the sample by SDS-PAGE overnight at 10 °C. (For optimal protein resolution, the gels must be long enough) |
| iv. | Stain the gel by an appropriate technique compatible with Mass Spectrometry. |
Analyze results
| i. | Compare bands between the lanes of gel between the experimental group and the control group, and identify the interactor referring to corresponding database. |
| ii. | Further confirm the protein-protein interaction by co-immunoprecipitation, Yeast-two Hybrid etc. |
In some steps, spin columns can be used to improve assay speed, protein recovery, reproducibility, and decrease elution volumes [2].
Question 1 (Q1): In MS analysis, many or all bands are identified as GST.
Possible cause: Very large amount of GST recombinant protein used as bait.
Possible solution: Cross-link the GST fusion protein to the GSH beads, or reduce the amount of GST fusion protein used.
Q2: Intense staining of four to five proteins of 20-30 KDa on SDS-PAGE gel, which interfere with the detection of other proteins.
Possible cause: Very large amount of endogenous tissue GST present in the tissue lysate, binding to the remaining free glutathione sites of the GSH beads.
Possible solution: Rigorously pre-clear lysate with GSH beads; if the GST fusion protein is cross-linked to the beads, wash the beads 2 to 3 times to specifically elute the GSTs.
Q3: No specific binding proteins are observed for the protein of interest relative to control.
Possible cause A: Bacterially produced GST fusion proteins are not correctly folded or post- translationally modified.
Possible solution A: Change expression system according to literature on protein of interest and on bacterial expression systems, or use a nonbacterial expression system, for example, in vitro translation, as long as sufficient GST fusion protein for pull-down can be obtained.
Possible cause B: Tissue source for lysate inappropriate for protein of interest.
Possible solution B: Change tissue source: use a tissue that is known to express the protein of interest, or screen several tissues and analyze simultaneously by SDS-PAGE, to compare binding proteins from different tissues; scale up chosen tissue by 10-fold; try tissues isolated from animals of different developmental stages.
Possible cause C: Washing or binding buffers may be too stringent (salt, detergent concentration interferes with binding).
Possible solution C: Dilute out salt or detergent further; homogenize pellets n smaller volumes, or use buffers for tissue extraction that have different properties.
Q4: Large amount of nonspecific proteins are eluted from GST fusion protein beads after pull-down.
Possible cause: Hydrophobic (largely membrane) proteins may adhere to the beads when washed without detergent.
Possible solution: Wash the beads with Triton X-100 containing wash buffer, then thoroughly wash the beads again with Triton X-100 free wash buffer.
Q5: There is still GST fusion protein eluted from the beads after chemical cross-linking, which interferes with pull-down experiment.
Possible cause A: The cross-linking reaction was incomplete or inefficient, or cross-linker has degraded.
Possible solution A: Ensure that sufficient cross-linker is used; requantitate the level of protein bound to GSH beads; cross-link for longer durations (up to 2 h); use fresh cross-linker; switch to more soluble cross-linker.
Possible cause B: GSH washing to remove small amount of non-cross-linked fusion protein was inadequate, or inefficient.
Possible solution B: Wash the column after cross-linking with bead recycling buffer, and allow the solution to pass through.
Q6: Interacting protein was not isolated.
Possible cause A: Weak or transient interactions.
Possible solution A: Decrease wash times or decrease the ion concentration of the buffer.
Possible cause B: Low expression level of prey protein.
Possible solution B: Increase the amount of prey protein.
[1] Coligan J E, Dunn B M, Ploegh H L, et al. Current protocols in protein science[M]. John Wiley & Sons Inc., New York, 2007.
[2] Brymora A, Cousin M A, Roufogails B D, et al. Enhanced protein recovery and reproducibility from pull-down assays and immunoprecipitations using spin columns[J]. Anal Biochem, 2001, 295(1): 119-22.
[3] Thery C, Amigorena S, Raposo G, et al. Current protocols in cell biology[M]. John Wiley, New York, 2006.
Fill out this form and one of our experts will respond to you within one business day.