Carbohydrates are involved in many important signaling pathways, and carbohydrate recognition is a key component of many biological processes including innate immunity, metabolism, and immune response. For example, glycans located on cell surfaces or secreted biomolecules play a crucial role in cell–cell interaction, including the interaction between host cell and pathogens. The physiological and pathological significance of glycan–protein interactions are drawing increasing attention in the field of structure-based drug design. However, it has been notoriously difficult to study these interactions by X-ray crystallography since the sugar moieties usually have a heterogeneous chemical structure and conformation. Computational molecular modeling techniques offer an attractive alternative method for the study of protein-carbohydrate recognition and specificity.
At Profacgen, we predict these protein–carbohydrate interactions through the combination of a genetic conformational search algorithm coupled with an empirical free energy function specific for carbohydrates. Our docking procedure allows for a systematic exploration of the orientations and positions of carbohydrate ligand into a protein cavity (which is usually more open and less well defined compared to small molecule binding sites), and assesses free energies of complex formation. Experiment-derived constraints can also be applied to the docking protocol to restrict the search space for the sugar as well as the protein molecule. The energy function is specifically designed for carbohydrates, which is used for the clustering of binding poses and exploration of the most energetically favorable complex structure. The relative stability of protein ligand complexes can be further evaluated by performing a short molecular dynamics simulation run.
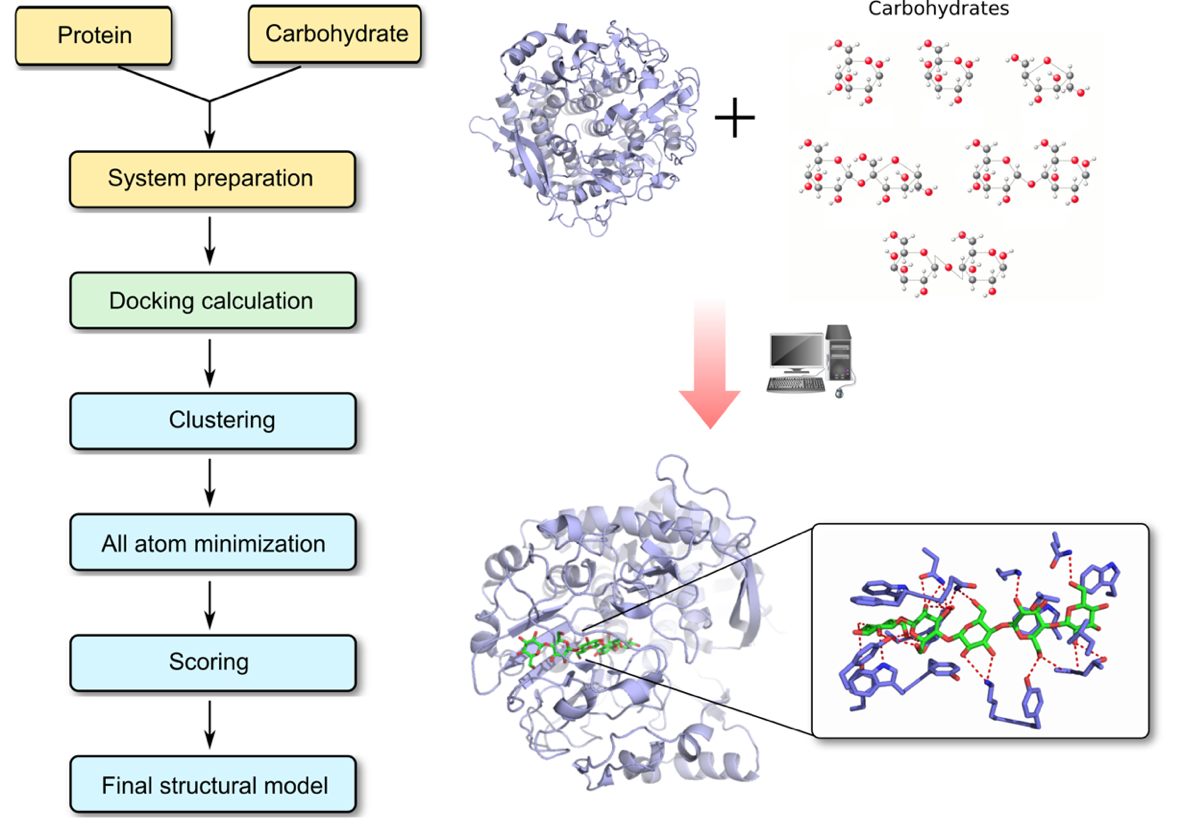
Given the growing importance of carbohydrate-binding proteins as potential drug targets, our computational modeling method is well suited to provide an atomic-level understanding of carbohydrate recognition which reveals some of the distinctive molecular features unique to protein-carbohydrate complexes. Our approach can predict the interaction between proteins and singe carbohydrate ligands, and it also has capability to predict the binding poses and binding affinities of glycans to glycan-binding proteins.
We provide the service in a customizable fashion to suit our customers’ specific research goals. Please do not hesitate to contact us for more details about our protein–carbohydrate docking service.
Fill out this form and one of our experts will respond to you within one business day.