Molecular dynamics (MD) simulation is a computer simulation technique that allows one to predict the time dependent evolution of a system of interacting particles using laws of physics. Studying the dynamic development of a biological system with the consideration of protein flexibility is of vital importance, since many biological phenomena involving proteins are very dynamic processes, which include transport, molecular recognition, enzyme catalysis, and allosteric regulation. The static models produced by NMR, X-ray crystallography, and homology modeling offer valuable insights into macromolecular structure, while MD simulation can provide atomic-level information about protein conformational changes and binding thermodynamics under predefined physiological conditions (e.g., temperature, pressure), allowing for the study of a protein system at a timescale that is not accessible by current experimental approaches.
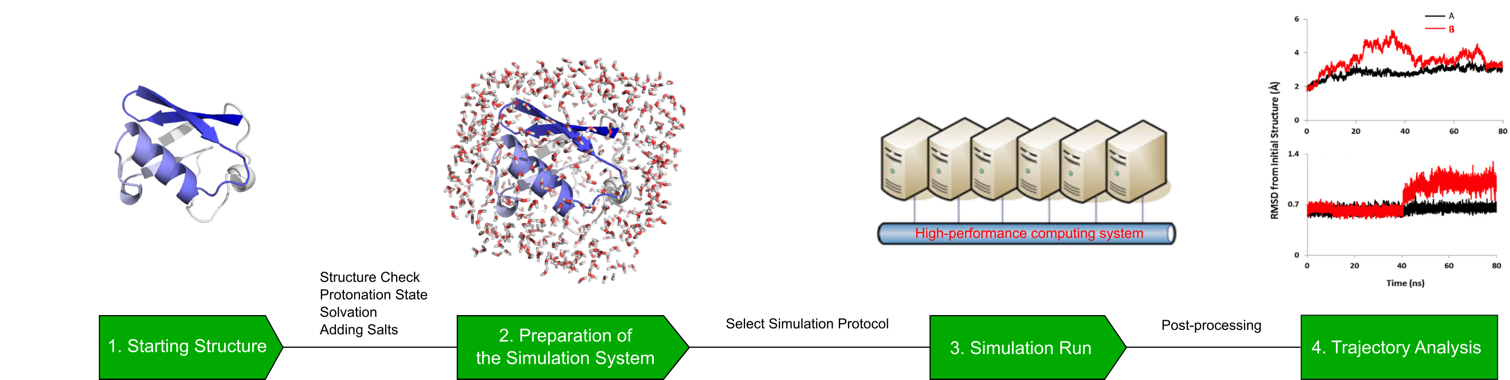
Profacgen studies molecular dynamics of protein systems using state-of-the-art software tools. The general workflow of our MD simulation service follows these steps: First, a model system is selected, in which missing segments are fixed and the protonation states determined. Then the system is energy minimized and equilibrated by solving Newton's equations of motion until the properties of the system no longer change with time. After equilibration, a production run is performed for an appropriate period of time to output trajectories, which are then analyzed for the properties of interest.
We take advantage of the MD simulation method to help customers predict the time dependent changes in a protein system, which can contain proteins, DNAs/RNAs, lipids and other small ligands, allowing for the exploration of events of biological and pharmaceutical importance. Specifically, simulation can be performed to characterize protein flexibility, refine experimentally determined structures, evaluate protein-ligand binding, study biocatalysis or even monitor the protein folding process. MD simulation is also particularly useful in computer-aided drug discovery for the identification of cryptic or allosteric binding sites, the enhancement of traditional virtual-screening methodologies, and the direct prediction of small-molecule binding energies.
We provide the service in a customizable fashion to suit our customers’ specific research goals. Please do not hesitate to contact us for more details about our molecular dynamics simulation service.
Fill out this form and one of our experts will respond to you within one business day.