Protein co-expression in Escherichia coli (E. coli) is a widely adopted and cost-effective strategy for producing multi-protein complexes required for biochemical, structural, and functional studies. By simultaneously expressing multiple interacting proteins within a single prokaryotic host, co-expression enables proper folding, enhanced solubility, and efficient assembly of biologically active complexes.
At Profacgen, we provide a comprehensive E. coli-based protein co-expression platform that integrates advanced vector engineering, multi-gene expression strategies, and optimized production workflows. Our services support the generation of di- and multi-subunit complexes, chaperone-assisted folding systems, and high-throughput expression screening. With extensive experience in both standard and complex co-expression systems, we deliver high-quality, functional protein assemblies tailored to diverse research and industrial applications.
Multi-protein complexes are involved in essentially all cellular processes, including transcription, translation, signaling pathways, and metabolic regulation. The function of a protein depends not only on its intrinsic physicochemical properties but also on its interaction status with binding partners and the characteristics of those partners.
In many cases, expressing a protein in isolation—especially in heterologous systems—results in insoluble, misfolded, or nonfunctional products. This is particularly true in prokaryotic expression systems, where the absence of cofactors, post-translational modifications, or interacting partners can limit proper folding and activity.
Protein co-expression addresses these limitations by enabling the simultaneous production of multiple proteins within the same host cell, allowing them to interact and assemble into functional complexes during biosynthesis.
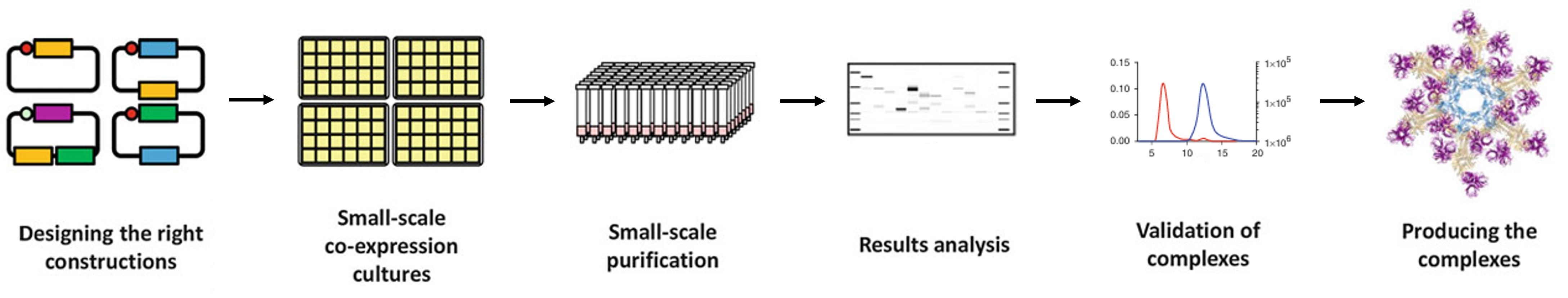 Figure 1. Pipeline of complex reconstitution by co- expression in Escherichia coli. (Adapted form Vincentelli and Romier, 2016)
Figure 1. Pipeline of complex reconstitution by co- expression in Escherichia coli. (Adapted form Vincentelli and Romier, 2016)
This approach provides several key advantages:
As a result, protein co-expression has become a recognized and powerful strategy for generating protein complexes in sufficient quantities for:
Among available systems, E. coli remains one of the most widely used hosts due to its rapid growth, ease of genetic manipulation, and cost-effectiveness. While more complex systems such as yeast, insect, or mammalian cells may be required for certain proteins, E. coli co-expression offers a highly efficient alternative for many applications.
Extensive studies and practical experience have demonstrated that co-expression in E. coli can successfully produce a wide range of functional protein complexes, making it a valuable tool for both academic and industrial research.
Profacgen provides a comprehensive and advanced E. coli protein co-expression platform, designed to support both standard and complex multi-protein expression projects. Our services are benchmarked against leading protein expression providers and incorporate flexible strategies to maximize success rates.

We provide multiple co-expression strategies to accommodate different protein systems:
Co-Expression from a Single Vector
This approach allows multiple genes to be carried on a single plasmid, ensuring coordinated expression.
Advantages:
Co-Expression from Multiple Vectors
Multiple plasmids are introduced into the same host cell to express different proteins.
To ensure plasmid compatibility and stability, we design vectors with:
Advantages:
We support a wide range of co-expression applications, including:
Profacgen also supports protein co-expression network analysis, enabling:
This service provides valuable support for systems biology and drug discovery research.
Client Requirements:
A research group aimed to produce a three-subunit enzyme complex for biochemical and structural characterization. Individual expression of each subunit in E. coli resulted in insoluble inclusion bodies and minimal enzymatic activity, preventing complex assembly and stalling the client's structural studies timeline.
Our Solution:
Profacgen designed a polycistronic single-vector system to co-express all three subunits simultaneously, ensuring proper stoichiometry. Chaperone co-expression was introduced to enhance folding of this challenging multi-protein complex. Expression conditions were systematically optimized, including low-temperature induction and varied media compositions, to promote solubility and correct assembly.
Final Results:
The optimized co-expression system yielded a soluble, correctly assembled enzyme complex with all three subunits present. Activity assays confirmed high enzymatic function comparable to native standards. The purified complex was successfully utilized for crystallization trials and structural determination, significantly accelerating the client's research progress.
Client Requirements:
A client required expression of a difficult therapeutic protein that consistently aggregated and formed inclusion bodies when expressed alone in E. coli, preventing downstream biochemical characterization and functional studies.
Our Solution:
We implemented a dual-vector co-expression system, expressing the target protein alongside a customized set of molecular chaperones and foldases including GroEL-GroES and DnaK-DnaJ-GrpE. Expression parameters including temperature, inducer concentration, and chaperone induction timing were carefully optimized to promote proper folding over aggregation.
Final Results:
The co-expression strategy dramatically improved protein solubility, with the majority of the target protein recovered in the soluble fraction. Purified material demonstrated correct folding by CD spectroscopy and retained full functional activity in biochemical assays. The client successfully advanced to interaction studies and structural characterization using the stabilized protein.
Consult Our Experts on Your Project
With a well-established platform for multi-protein co-expression in E. coli, Profacgen enables efficient production of soluble and functional protein complexes for a wide range of applications. Whether your project involves simple heterodimers or more complex assemblies, our team can design and optimize a strategy tailored to your specific requirements.
Contact us today to discuss your protein co-expression project and receive a customized solution for high-yield and high-quality complex production.
For projects that do not require multi-protein assembly, single-protein expression in E. coli may offer a more streamlined and cost-effective solution. Our bacterial expression platform supports high-yield production with rapid turnaround and scalable processes.
References:
Fill out this form and one of our experts will respond to you within one business day.