Pichia pastoris is a methylotrophic yeast and can be used as a heterologous expression system [1]. It is a single-celled microorganism that is easy to manipulate and culture as Escherichia coli [2]. In the meantime, as a eukaryote, it possesses the capabilities of post-translational modifications performed by higher eukaryotic cells. Proteins that express as inactive inclusion bodies in bacterial expression systems can therefore be produced as biologically active molecules in P. pastoris, with correct proteolytic processing, folding, disulfide bond formation and glycosylation. Besides, the P. pastoris system is also regarded as being easier, faster and more cost-efficient to use than expression systems developed from higher eukaryotes such as insect cells and mammalian tissue culture cell systems. The protein expression levels in P. pastoris system are also generally much higher. As yeast, it shares the advantages of easy genetic manipulations with Saccharomyces; moreover, it possesses the advantage of 10-100 folds higher heterologous protein expression levels. All the above features have made Pichia a very useful protein expression system. It is worth mentioning that, several techniques developed for Saccharomyces initially can also be applied to Pichia, including transformation by complementation, gene disruption and replacement.
Pichia pastoris is a type of methylotrophic yeast and thus is capable of metabolizing methanol as its sole carbon source [3]. The first step in the metabolism of methanol is the oxidation of methanol into formaldehyde by molecular oxygen. The reaction also generates hydrogen peroxide except for formaldehyde. To avoid the toxicity of hydrogen peroxide, methanol metabolism normally takes place within the peroxisome, a specialized cellular organelle. Alcohol oxidase, the enzyme that catalyzes the oxidation of methanol has a poor affinity for oxygen, and thus Pichia pastoris has to generate large amounts of the enzyme as compensation. In the meantime, the promoter regulating the production of the enzyme is therefore used to drive heterologous protein expression in Pichia.
There are two genes code for alcohol oxidase in Pichia, AOX1 and AOX2 [4]. Generally, a majority of the activity of the enzymes relies on the AOX1 gene. Its expression is largely controlled by methanol. A plasmid-born version of the AOX1 promoter is applied to induce expression of inserted gene of interest for the desired heterologous protein [3,4]. Although AOX2 is about 97% homologous to AOX1, its growth on methanol is much slower than that of AOX1.
To be more specific, the expression of the AOX1 gene is controlled at the level of transcription. The regulation of AOX1 gene is a two-step process, which includes a repression mechanism and an induction mechanism. Since growth on glucose can repress transcription process even when methanol exists; growth on glycerol is recommended for optimal induction with methanol.
As to the expression pattern of heterologous protein in Pichia pastoris, it can be either intracellular or secreted, depending on the chosen expression vector. Secretion requires the presence of a signal peptide on the protein via secretory pathway. A major advantage of secreted expression of heterologous protein is that Pichia pastoris itself secrets very low levels of native proteins. Thus the secreted heterologous comprises the majority of the total protein in Pichia growth medium and therefore simplifies subsequent purification procedure of the protein [1].
Typically, posttranslational modifications in Pichia include removal of signal sequences, folding, disulfide bridge formation and O-/N-linked glycosylation. In mammals, O-linked oligosaccharides are composed of sugars including N-acetygalactosamine, galactose and sialic acid [2]. While in lower eukaryotes like P.pastoris, the added O-linked oligosaccharide simply refers to mannose residues.
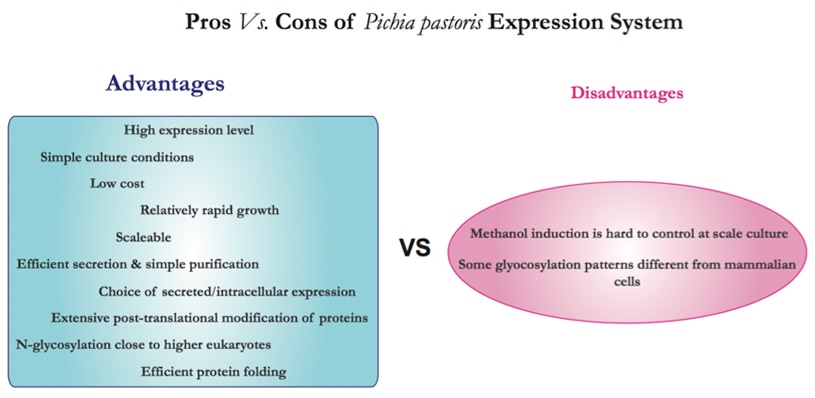
P. pastoris has become a highly successful system for the expression of heterologous genes. Several important factors that contribute to its rapid spread and wide range application are summarized as following points [2].
a.A promoter derived from the alcohol oxidase I (AOX1) gene of P.pastoris that is appropriate for the precise expression of foreign genes.
b. Similar techniques of the molecular genetic manipulation in P.pastoris to those from Saccharomyces cerevisiae, one of the most well-characterized experimental systems in modern biology.
c. P.pastoris prefers respiratory growth, which is a key physiological trait which allows its culturing at high cell densities relative to fermentative yeasts.
Our trained custom service group will perform all the steps for the fastest expression of your target protein using the P.pastoris Expression System, which will definitely save your resources and time.
The flow chart below illustrates the overall experimental process required for recombinant protein production using the Pichia Expression System.
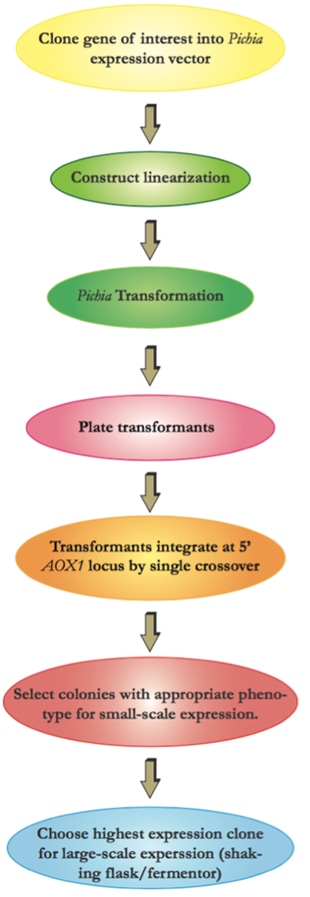
The following protocol can be a reference for you to understand more details of the techniques utilized in our lab and workflow involved in producing the protein of interest using P.pastoris.
1. Select the appropriate Expression Vector for your target gene.
You may choose to express your protein intracellularly if your protein is cytosolic and non-glycosylated. You can also have your target protein secreted into the culture media if your protein is generally secreted, glycosylated or directed to an intracellular organelle.
2. Transformation into E.coli.
| i. | Insert the target gene in frame with the expression vector. |
| ii. | Transform the E.coli with the ligation mixture by electroporation or chemical methods. |
| iii. | Add LB medium to the cells for recovery after heat shock or electroporation. |
| iv. | Plate on LB medium with Bleomycin. |
| v. | Incubate overnight. |
3. Analyze transformants.
| i. | Select Bleomycin-resistant colonies and inoculate into LB medium with Bleomycin. |
| ii. | Grow overnight and isolate plasmid DNA. |
| iii. | Sequence the gene construct to confirm the correct insertion of gene within the vector. |
4. Preparation for transformation
Prior to transformation and selection in Pichia, the plasmid should be linearized. Vector linearized within the 5’ AOX1 region will integrate into the host 5’ AOX1 region by gene insertion.
| i. | Digest the plasmid DNA with selected restriction enzymes. |
| ii. | Check the digested mixture for complete linearization by agarose gel electrophoresis. |
| iii. | Once the vector is confirmed to be completely linearized, use heat treatment for inactivation or add EDTA to cease the reaction immediately. |
| iv. | Use phenol/chloroform for extraction and ethanol for precipitation of the DNA. |
| v. | Centrifuge the solution and wash the pellet DNA with ethanol. |
| vi. | Resuspend the DNA in sterile water after air-dry for immediate use or store at -20°C. |
5. Electroporation of Pichia.
The method of electroporation is strongly recommended because it gives the highest transformation frequency in Pichia.
| i. | Grow the selected P.pastoris strain in yeast medium overnight. |
| ii. | Add fresh medium in the overnight culture. Grow overnight again. |
| iii. | Centrifuge the cells and resuspend with ice-cold, sterile water. Repeat twice. |
| iv. | Centrifuge the cells, then resuspend with ice-cold sorbitol. Repeat twice. Place the cells on ice before use. |
| v. | Mix the cells with linearized DNA. |
| vi. | Transfer the mixture to an ice-cold electroporation cuvette. Incubate on ice. |
| vii. | Add ice-cold sorbitol to the cells, then transfer the cuvette content and incubate at 30°C. |
| viii. | Spread the cells on plates for Bleomycin selection. |
| ix. | Incubate the plates until colonies form. |
| x. | Pick 10-20 colonies for further selection. |
6. Analyze Pichia transformants
| i. | Inoculate yeast medium with the chosen single colony of Pichia strain. Grow overnight. |
| ii. | Dilute the cells from the overnight culture and grow approximately 4-6 h. |
| iii. | Centrifuge the cells and keep the pellet. |
| iv. | Resuspend the cell pellet and get the competent cells. |
| v. | The cells can be kept at room temperature and used for transformation assay at once or frozen for storage. |
7. Transformation assay
The following procedure provides details of transformation of freshly prepared or frozen competent Pichia cells. Note that transformation efficiency may vary among different Pichia strains and expression vectors used.
| i. | Add linearized recombinant vector DNA to the competent cells. |
| ii. | Add solution containing PEG into the DNA/cell mixture. |
| iii. | Incubate the transformation reaction for 1h at 30°C. Mix the reaction solution to increase transformation efficiency. |
| iv. | Treat with heat shock and split the cells into microcentrifuge tubes for incubation. |
| v. | Centrifuge the cells and keep the pellet. |
| vi. | Resuspend and combine the cells. |
| vii. | Plate the cells for selection. Incubate the plates until colonies appear. |
8. Determine the Mut (Methanol Utilization Slow) phenotype.
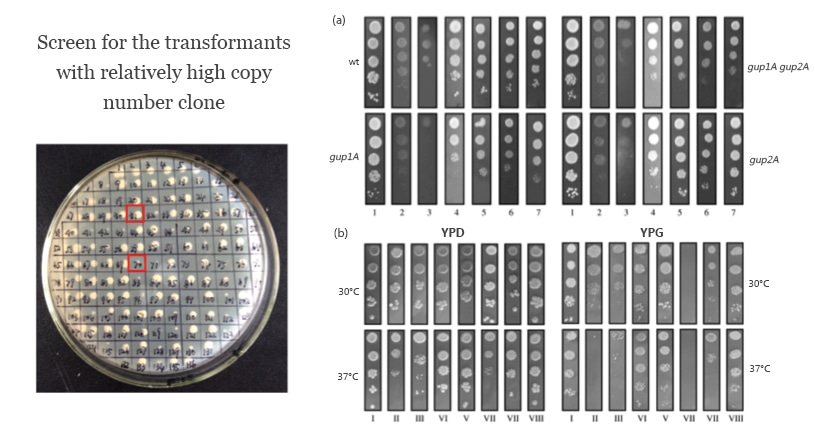
Identify the expression levels among several chosen Mut phenotypes by SDS-PAGE analysis and screen for the high-expression Pichia recombinant clones. This may help optimize the expression condition of the recombinant clone.
The effectiveness of expression conditions of Pichia strain with Mut phenotype can be tested as follows.
| i. | Include a control gene transformed with the parent vector as a control for background intracellular expression. |
| ii. | Inoculate a single colony in a baffled shaking flask and grow at 28-30°C. |
| iii. | Harvest the cells and centrifuge at room temperature. |
| iv. | Discard supernatant and resuspend cell pellet. Cover the flask with 2 layers of sterile gauze and continue growth. |
| v. | Add methanol every 24 hours to maintain induction. |
| vi. | Transfer 1 ml of the expression culture to a fresh microcentrifuge tube to analyze expression levels at a series of time points (i.e., 0-96 h post-induction). |
| vii. | Determine the optimal time to harvest the cells. Centrifuge the cells at room temperature. |
| viii. | For secreted expression, transfer the supernatant to a fresh tube and store until ready to assay. While, for intracellular expression, discard the supernatant and store the cell pellets until ready to assay. |
| ix. | Analyze the supernatants and cell pellets respectively for protein expression by SDS-PAGE and Western blot. |
The above steps can be adjusted to optimize expression for your target protein.

9. Scale-up expression.
Once expression condition is optimized, larger baffled flasks or fermentation will be utilized to increase the culture volume for scale-up production of target protein.
Note that proteins secreted into the media are normally more than 50% homogeneous and need additional purification. Therefore it is an optimal step to concentrate the protein before the purification process if the expression level is low. Commonly used methods for this step are ammonium sulfate precipitation, lyophilization and centrifuge concentrator for small volumes [5].
Click here to contact us for more technical information.
[1] Higgins D R. Overview of protein expression in Pichia pastoris[J]. Curr Protoc Protein Sci, 2001, Chapter 5: Unit5 7.
[2] Higgins D R, Cregg J M. Introduction to Pichia pastoris[J]. Methods Mol Biol, 1998, 103: 1-15.
[3] Lundblad R L. Glycosylation in Pichia pastoris[J]. Biotechnol Appl Biochem, 1999, 30 ( Pt 3): 191-2.
[4] Bretthauer R K, Castellino F J. Glycosylation of Pichia pastoris-derived proteins[J]. Biotechnol Appl Biochem, 1999, 30 ( Pt 3): 193-200.
[5] Buckholz R G, Gleeson M A. Yeast systems for the commercial production of heterologous proteins[J]. Biotechnology (N Y), 1991, 9(11): 1067-72.
Fill out this form and one of our experts will respond to you within one business day.