Protein degradation and stability provides the ultimate regulation of protein functions. The traditional 35S-based pulse-chase degradation measurement method requires the use of a radioactive isotope, which is of restricted access and is unfavorable in many laboratories. As an advanced one-stop protein degradation service platform, Profacgen offers multiple technical services for degradation assay and stability analysis.
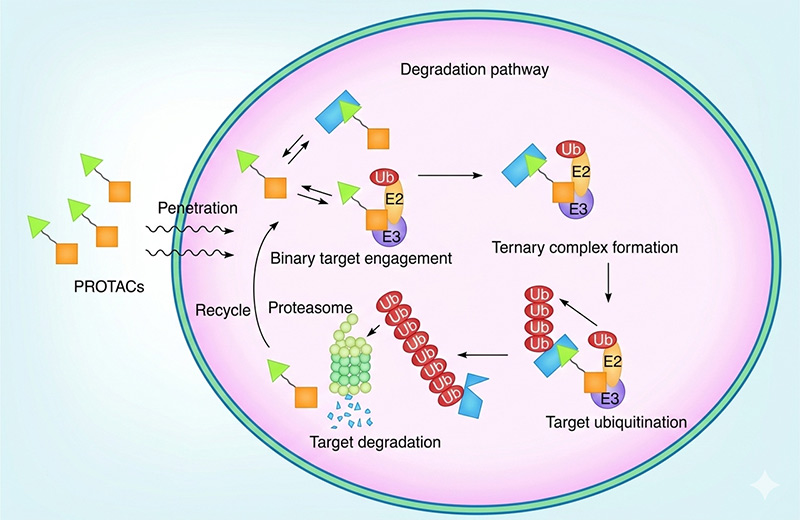
Figure 1. Degradation pathway for proteolysis targeting chimeras
Antibody based Western blot or ELISA assay
Western blot or ELISA detection rely on the availability of good antibodies to the target of interest or the use of phenotypic endpoints. As a classic method, they are widely used in in vitro degradation assay and stability analysis, the only regret is that the efficiency is dependent on the pull-down effect by antibody.
In eukaryotic cells, proteasomes and lysosomes are two main organelles, in which proteins are degraded into peptides and amino acids. This method studies the protein degradation on a proteome-wide scale by detection of the intermediate peptides produced from the intracellular degradation of proteins using sequencing-based tandem MS.
Real-time degradation kinetics measurement
This assay shows comprehensive real-time degradation and recovery profiles for each target, precisely quantifying degradation rates, maximal levels of degradation (Dmax), and time frame. Profacgen combines endogenous tagging and luminescent technology to kinetically measure target protein levels with high precision and without disruption of native expression levels or transcriptional regulation. Coupling with optimized bioluminescence resonance energy transfer method allows for kinetic measurement of intracellular protein interaction, such as ternary complex formation, ubiquitination, and protein degradation-target engagement.
Enzyme Fragment Complementation (EFC) technology
By combining CRISPR technique with EFC technology, Profacgen has biosensor lines that allows sensitive quantitation of protein degradation and stability using a homogeneous assay format. This technology takes advantage of the ubiquitin-proteasome protein degradation system to measure protein turnover and identify new molecular entities that modulate protein levels for small molecule therapeutic development.
Time-resolved fluorescence resonance energy transfer (TR-FRET) technique
TR-FRET detection is a well-established and robust technology frequently used in drug screening. The principle is based on protein quantification after small-molecule compound-induced stabilization. It enables highly sensitive detection of protein kinetics using affinity-tagged protein and owns the advantages of a time-resolved measurement.
The workflow of our services:
1. Identify demand and project design.
2. Sample submission and quality control.
3. Measurement and progress communication.
4. Data analysis and report delivery.
Profacgen promises to work closely with clients, please contact us for more information.
1. Zhang Y, Jiang Y, Wang S. (2010). Development of an Enzyme-Linked Immunosorbent Assay To Detect Benzylpenicilloic Acid, a Degradation Product of Penicillin G in Adulterated Milk. Journal of Agricultural & Food Chemistry, 58(14), 8171–8175. doi: 10.1021/jf101243z
2. Wu P, Lu M, Cui X, Yang H, Yu S, Zhu J, Sun X, Lu B. (2016). A high-throughput-compatible assay to measure the degradation of endogenous Huntingtin proteins. Acta Pharmacologica Sinica, 37(10), 1307–1314. doi:10.1038/aps.2016.31
3. Shen Y, Hixson KK, Tolić N, Camp DG, Purvine SO, Moore RJ, Smith RD. (2008). Mass Spectrometry Analysis of Proteome-Wide Proteolytic Post-Translational Degradation of Proteins. Analytical Chemistry, 80(15), 5819–5828. doi:10.1021/ac800077w
4. Riching KM, Mahan S, Corona C, McDougall MG, Vasta JD, Robers MB, Urh M, Daniels DL. (2018). Quantitative live-cell kinetic degradation and mechanistic profiling of PROTAC mode of action. ACS Chemical Biology, 13, 2758-2770. doi:10.1021/acschembio.8b00692
5. Schulze J, Moosmayer D, Weiske J, Fernández-Montalván A, Herbst C, Jung M, Haendler B, Bader B. (2015). Cell-Based Protein Stabilization Assays for the Detection of Interactions between Small-Molecule Inhibitors and BRD4. Journal of Biomolecular Screening, 20(2), 180–189. doi:10.1177/1087057114552398
6. Xingui Liu, Xuan Zhang et al. Assays and technologies for developing proteolysis targeting chimera degraders. Future Med Chem. 2020 Mar; 12(12): 1155–1179. doi: 10.4155/fmc-2020-0073
Fill out this form and one of our experts will respond to you within one business day.