Disulfide bonds are covalent linkages between cysteine residues that stabilize protein tertiary and quaternary structures, govern folding fidelity, and maintain biological activity. Profacgen's Disulfide Bond Mapping services employ non-reducing peptide mapping coupled with high-resolution LC-MS/MS, partial reduction strategies, and free-thiol quantification to deliver unambiguous assignment of intra- and interchain disulfide connectivity. Whether confirming the canonical architecture of a monoclonal antibody, resolving the complex hinge-region isoforms of IgG2, or detecting mispaired disulfides in a novel fusion protein, our multi-method platform ensures comprehensive structural verification with regulatory-grade documentation.
Correct disulfide bond pairing is a critical quality attribute (CQA) for all cysteine-containing biotherapeutics. Mispaired, scrambled, or free-thiol-containing variants can compromise structural integrity, reduce potency, increase aggregation propensity, and elevate immunogenicity risk. Regulatory agencies explicitly mandate disulfide bond characterization under ICH Q6B guidelines, requiring determination of the number and positions of free sulfhydryl groups and disulfide bridges to the extent possible.
Profacgen addresses these requirements through a multi-tiered analytical strategy. Our foundational approach compares non-reducing and reducing peptide maps by LC-MS/MS to identify disulfide-linked peptides and confirm expected connectivity. For densely cysteine-rich regions or complex interchain patterns—as seen in IgG2 hinge isoforms and bispecific antibodies—we deploy partial reduction coupled with cyanylation-induced cleavage or electron transfer dissociation (ETD) to resolve ambiguous linkages. Free-thiol content is quantified by the Ellman assay or fluorescent maleimide labeling to complement the mapping data. Together, these methods provide orthogonal evidence of correct folding and enable detection of trace-level mispairing.
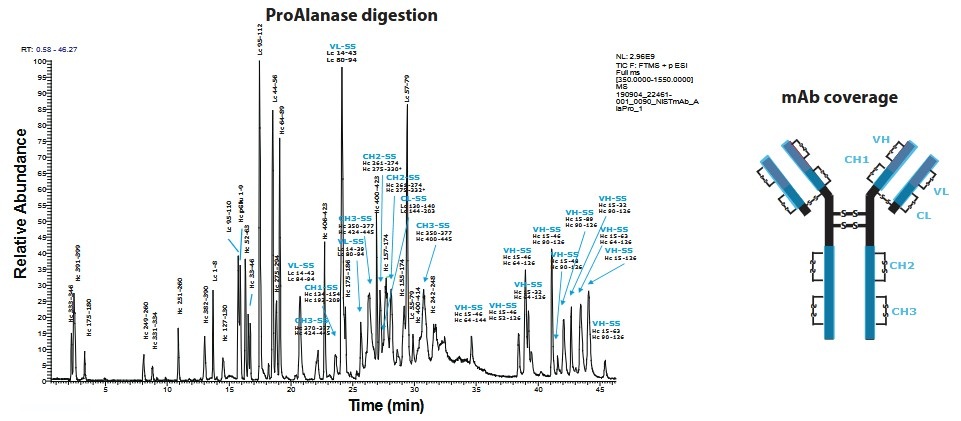 Figure 1. Example of disulfide bond mapping for antibodies. (Samodova et al., 2020)
Figure 1. Example of disulfide bond mapping for antibodies. (Samodova et al., 2020)
Profacgen provides end-to-end disulfide bond characterization tailored to discovery, development, and quality control applications. Our offerings include:
| Service Component | Description |
|---|---|
| Non-Reducing Peptide Mapping & LC-MS/MS |
|
| Partial Reduction & Cyanylation-Induced Cleavage |
|
| Free Sulfhydryl (Thiol) Quantification |
|
| IgG2 Hinge Isoform & Bispecific Characterization |
|

Background:
A biopharmaceutical company observed an increase in high-molecular-weight species (HMWS) and a decrease in potency during accelerated stability testing of their IgG1 monoclonal antibody at 40 °C. Because disulfide scrambling is a known degradation pathway that can generate covalent aggregates and alter antigen-binding affinity, the client needed to determine whether the stress conditions had induced non-native disulfide pairing.
Our Solution:
Profacgen performed parallel non-reducing peptide mapping on control and heat-stressed samples using an optimized one-pot digestion protocol with 8 M urea plus guanidine-HCl at 50 °C, followed by two-step trypsin/Lys-C digestion. Free cysteines were alkylated with iodoacetamide prior to denaturation to prevent scrambling artifacts. Disulfide-linked peptides were identified by comparing non-reduced and reduced LC-MS/MS data. Extracted ion chromatograms of expected disulfide-bridged peptides were quantified across both conditions.
Final Results:
The stressed sample exhibited a 12-fold increase in a non-native disulfide linkage between Cys214 (heavy chain) and Cys226 (light chain), confirming heat-induced scrambling in the hinge region. Native interchain bonds remained intact at >95 %, but the scrambled population correlated directly with the HMWS increase. The client used these data to establish a maximum allowable temperature excursion limit of 25 °C for shipping, and Profacgen's report supported a successful regulatory response to the regulatory agency regarding the observed stability trend.
Background:
A biotechnology company developed an IgG2-based agonistic antibody intended to cross-link cell-surface receptors. The IgG2 subclass is unique in displaying three distinct hinge-region disulfide isoforms—A, A/B, and B—each with different inter-heavy-chain connectivity that affects receptor clustering activity. The client needed to quantify isoform distribution and confirm that their manufacturing process consistently produced the desired pseudo-isoform B structure stabilized by noncovalent interactions.
Our Solution:
We employed native cation-exchange chromatography–mass spectrometry (CEX-MS) using volatile salts to separate the intact isoforms without denaturation. To localize hinge-region linkages, we performed IdeS digestion to generate F(ab')2 fragments, which were then analyzed by non-reducing peptide mapping with an isotope-envelope confidence score for unambiguous identification of hinge-related peptides. Site-directed mutagenesis of cysteine residues and controlled redox treatment confirmed the elution order and linkage pattern of each isoform.
Final Results:
CEX-MS resolved three baseline-separated peaks corresponding to isoforms A (18 %), A/B (34 %), and B (48 %). The dominant isoform B population matched the expected noncovalent-stabilized structure required for optimal agonistic activity. Non-reduced peptide mapping confirmed the characteristic Cys232–Cys232 inter-heavy-chain linkage of isoform B with >99 % confidence. The client incorporated Profacgen's isoform quantitation into their release specifications, ensuring consistent clinical performance across manufacturing campaigns.
Background:
An oncology biotech company engineered a cysteine residue into the antibody constant region to enable site-specific conjugation of a cytotoxic payload. During process development, they observed variable drug-to-antibody ratios (DAR) and suspected incomplete oxidation of the engineered cysteine, leaving residual free thiols that competed with the conjugation chemistry. Accurate quantitation of free-thiol content was essential to optimize oxidation conditions and ensure batch-to-batch DAR consistency.
Our Solution:
Profacgen implemented a dual analytical approach. First, the Ellman assay (DTNB) was performed on intact antibody samples to measure total free-thiol content spectrophotometrically at 412 nm. Second, non-reducing peptide mapping with IAM alkylation was used to localize the free cysteine to the engineered site in the CH2 domain. Parallel intact mass analysis under non-reducing and reducing conditions confirmed the mass shift associated with thiol vs. disulfide states. The combined data provided both quantitative and site-specific evidence of cysteine oxidation efficiency.
Final Results:
Ellman quantitation revealed 0.42 free thiols per antibody molecule (target: <0.25), indicating suboptimal oxidation. Peptide mapping localized the free thiol exclusively to the engineered Cys239 position, ruling out native cysteine reduction as the source. The client adjusted the oxidation buffer pH from 7.4 to 8.2 and extended the oxidation time from 2 h to 4 h, reducing free-thiol content to 0.08 and stabilizing DAR at 2.02 ± 0.05. Profacgen's validated free-thiol method was transferred to the client's QC laboratory for routine lot-release testing.
Consult Our Experts on Your Project
References:
Fill out this form and one of our experts will respond to you within one business day.