Peptide mapping by LC-MS/MS is the cornerstone of biotherapeutic protein characterization, providing a site-specific molecular fingerprint that confirms primary sequence identity and monitors critical quality attributes (CQAs). At Profacgen, our Peptide Mapping (LC-MS/MS) services combine optimized enzymatic digestion workflows with high-resolution accurate-mass (HRAM) mass spectrometry to deliver comprehensive sequence coverage, precise post-translational modification (PTM) localization, and rigorous batch-to-batch comparability data. From monoclonal antibodies and fusion proteins to novel bioconjugates, our platform supports every stage of biopharmaceutical development, ensuring regulatory-compliant structural verification for IND, BLA, and biosimilar submissions.
Biopharmaceuticals are inherently complex, and their safety and efficacy depend on an unambiguous primary structure and tightly controlled post-translational modification profile. Regulatory agencies explicitly require peptide mapping as a fundamental characterization tool under ICH Q6B guidelines to confirm amino acid sequence identity, assess purity, and detect process- or storage-related degradation. Unlike intact mass analysis, which provides global mass confirmation only, peptide mapping resolves structural information to the single-residue level, enabling precise localization of modifications such as oxidation, deamidation, glycosylation, and sequence variants.
Profacgen's peptide mapping platform addresses these demands through a multi-attribute analytical strategy. We employ optimized reduction and alkylation protocols followed by single- or multi-enzyme digestion (trypsin, chymotrypsin, AspN, Glu-C, or Lys-C) to achieve >95 % sequence coverage even for challenging hydrophobic domains. Peptides are separated on advanced reversed-phase columns (C18 or C4) and analyzed by high-resolution LC-MS/MS using Orbitrap or Q-TOF platforms. This approach simultaneously confirms sequence identity, quantifies PTMs, monitors disulfide bond integrity, and supports biosimilarity assessments with unmatched sensitivity and reproducibility.
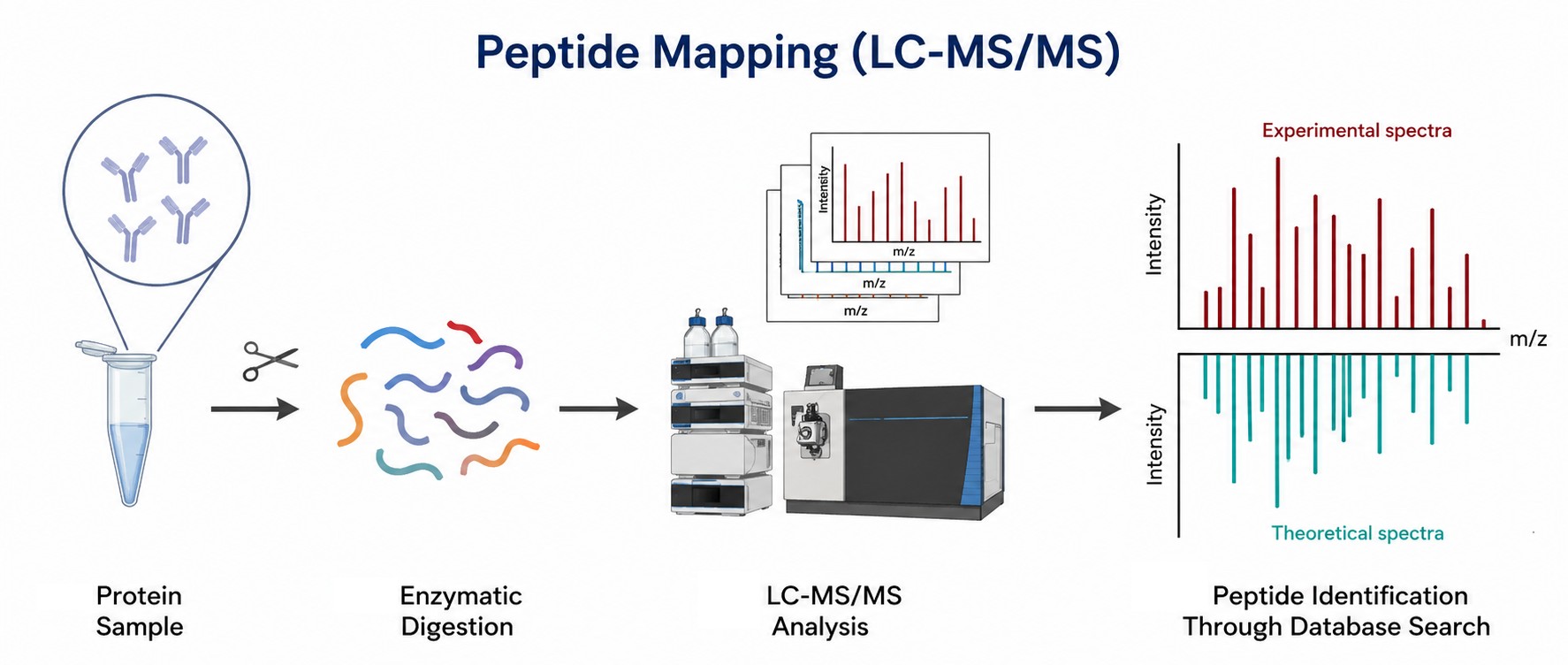 Figure 1. Representative peptide mapping workflow combining multi-enzyme digestion, reversed-phase separation, and high-resolution tandem mass spectrometry for comprehensive protein characterization.
Figure 1. Representative peptide mapping workflow combining multi-enzyme digestion, reversed-phase separation, and high-resolution tandem mass spectrometry for comprehensive protein characterization.
Profacgen provides end-to-end peptide mapping solutions tailored to discovery, development, and quality control applications. Our offerings include:
| Service Component | Description |
|---|---|
| Tryptic Peptide Mapping & Sequence Confirmation |
|
| Multi-Enzyme Digestion for Enhanced Coverage |
|
| PTM Detection & Quantification |
|
| Biosimilarity & Comparability Studies |
|
| Stability & Forced Degradation Monitoring |
|

Background:
A biopharmaceutical company observed an upward drift in acidic charge variants during cation-exchange chromatography of their Phase II monoclonal antibody over six months of stability storage. Because charge heterogeneity can impact pharmacokinetics and immunogenicity, the client needed to pinpoint the exact modification sites and quantify the rate of change to set shelf-life specifications and justify formulation adjustments.
Our Solution:
Profacgen performed reduced and alkylated tryptic peptide mapping under both native and stressed conditions. Using a Q-Exactive HF-X Orbitrap with high-energy collisional dissociation (HCD), we achieved 98 % sequence coverage. Comparative analysis of stressed versus control samples revealed a consistent +0.984 Da mass shift localized to an asparagine residue within the heavy-chain complementarity-determining region 2 (CDR-H2). Parallel AspN digestion confirmed the site, and extracted ion chromatograms enabled relative quantitation of deamidation across all stability time points.
Final Results:
The deamidation level at the CDRH2 asparagine increased from 3.2 % at t=0 to 18.7 % at 40 °C/75 % RH over three months. This rate exceeded the client's quality target, prompting a switch from phosphate-buffered to histidine-buffered formulation, which reduced the degradation rate by 60 %. Profacgen's data package, including raw MS files, peak area ratios, and validated method parameters.
Background:
A biosimilar developer required a comprehensive peptide mapping comparability package to demonstrate analytical similarity between their trastuzumab biosimilar candidate and the EU-sourced reference product. The study was critical for the totality-of-evidence approach mandated by EMA and FDA biosimilar guidelines, covering primary sequence, PTM profiles, and glycosylation site occupancy.
Our Solution:
We executed a multi-enzyme peptide mapping strategy combining trypsin, chymotrypsin, and AspN digestions to maximize sequence coverage across the F(ab')2 and Fc regions. Each digest was analyzed by reversed-phase nanoLC-MS/MS on an Orbitrap Exploris 480. PTM quantification included methionine oxidation, asparagine deamidation, N-terminal cyclization, and C-terminal lysine heterogeneity. Glycopeptides were identified using HCD-triggered electron transfer dissociation (ETD) to preserve glycan attachments.
Final Results:
Sequence coverage exceeded 99 % for both products, with 100 % peptide overlap and no detectable sequence variants. PTM profiles were statistically equivalent: methionine oxidation (2.1 % vs. 2.3 %), deamidation (4.8 % vs. 5.1 %), and C-terminal lysine variants (major: 62 % vs. 59 %). Glycosylation site occupancy at Asn300 was complete in both samples. The client submitted Profacgen's comparability report directly to regulatory authorities, accelerating their biosimilar approval timeline by eight months.
Background:
A gene-therapy company detected an anomalous peak in the peptide map of their recombinant Fc-fusion protein during lot-release testing. The peak was absent in the reference standard and represented approximately 0.8 % of the total peptide area. The client needed to determine whether the anomaly indicated a sequence variant, a process-related modification, or a contaminant—each with very different implications for product quality and patient safety.
Our Solution:
Profacgen isolated the anomalous peptide by preparative-scale reversed-phase fractionation and subjected it to high-resolution MS/MS analysis. The intact mass of the peptide was 128.1 Da higher than the expected tryptic peptide from the known sequence. Fragment ion mapping localized the mass shift to a single amino acid position where leucine was encoded. Database-independent de novo sequencing of the MS/MS spectrum unambiguously assigned the additional mass to an arginine residue substituting for leucine—a single-nucleotide polymorphism (SNP) in the expression construct.
Final Results:
The arginine-for-leucine substitution introduced a new tryptic cleavage site, explaining the unexpected peptide. The variant was traced to a low-frequency mutation in the stable cell line that had evaded earlier sequencing. The client used Profacgen's findings to implement clonal screening and genetic stability testing, eliminating the variant from subsequent manufacturing campaigns. The rapid 12-day turnaround prevented a costly batch rejection and reinforced the client's quality management system.
Consult Our Experts on Your Project
Fill out this form and one of our experts will respond to you within one business day.