Post-Translational Modification (PTM) Characterization
Profacgen's Post-Translational Modification (PTM) Characterization service provides comprehensive identification, localization, and quantification of covalent protein modifications, delivering the molecular evidence required to ensure product quality, demonstrate consistency, and satisfy regulatory expectations for biologic therapeutics.
Post-translational modifications—including glycosylation, oxidation, deamidation, disulfide bond formation, phosphorylation, and acetylation—profoundly influence protein activity, stability, half-life, and immunogenicity. These modifications are not genetically encoded and can vary with cell line, culture conditions, purification process, and storage history. Rigorous PTM characterization is therefore essential for defining critical quality attributes (CQAs), establishing release specifications, and supporting IND, BLA, and biosimilar submissions.
Background: PTM Characterization
PTMs represent a major source of molecular heterogeneity in biologics. Glycosylation patterns affect antibody-dependent cellular cytotoxicity (ADCC) and complement-dependent cytotoxicity (CDC); oxidation and deamidation can reduce potency and accelerate degradation; disulfide bond scrambling may compromise structural integrity; and phosphorylation or acetylation can modulate signaling activity. Regulatory agencies require thorough PTM profiling to demonstrate that a product is well-characterized and that manufacturing processes consistently yield the intended modification profile.
Profacgen's PTM platform addresses these requirements through orthogonal analytical techniques that localize modifications to specific residues, quantify their abundance, and assess their impact on product quality. Our integrated approach combines high-resolution mass spectrometry, chromatographic separation, and electrophoretic methods to deliver unambiguous, regulatory-compliant data packages.
- Glycosylation profiling and site-specific glycan analysis by HILIC and LC-MS/MS
- Oxidation, deamidation, and isomerization detection with site-specific localization
- Disulfide bond mapping, scrambling assessment, and free thiol quantification
- Phosphorylation, acetylation, methylation, and low-frequency modification screening
These analyses establish a comprehensive PTM fingerprint that supports process understanding, comparability assessment, and regulatory filing.
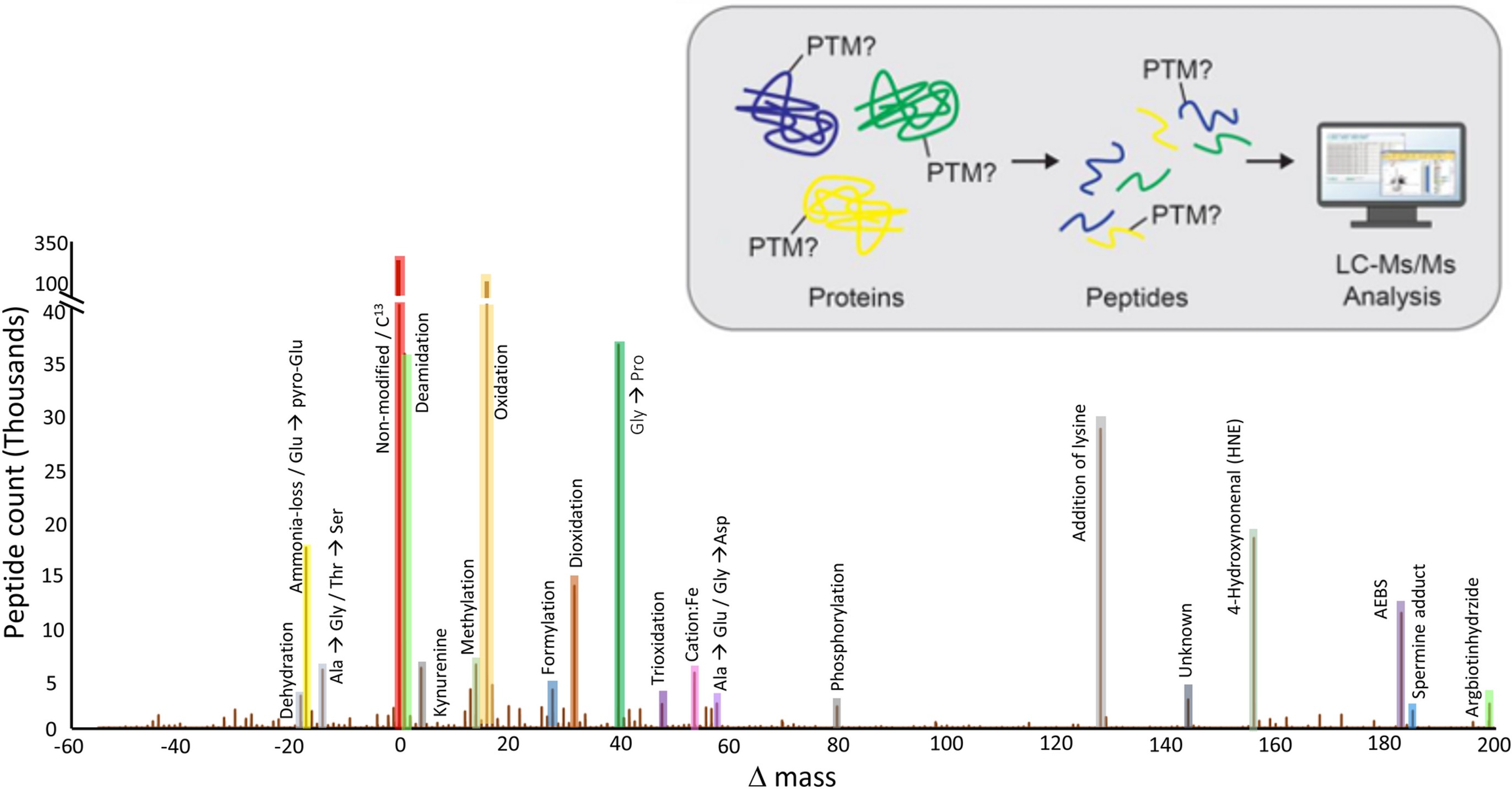 Figure 1. An example of PTM identifications in human hearts. (Adapted from Bagwan et al., 2021)
Figure 1. An example of PTM identifications in human hearts. (Adapted from Bagwan et al., 2021)
What We Offer: PTM Characterization Services
Our PTM Characterization platform integrates orthogonal analytical methodologies to deliver definitive, stage-appropriate modification evidence. We tailor technique selection and reporting depth to your regulatory pathway, from early clone screening through formal release testing, stability protocols, and biosimilar comparability studies.
Our PTM platform offers two dedicated analytical modules, each optimized for a specific modification class with tailored methodologies and interpretive frameworks.
Glycosylation Profiling
Comprehensive glycan analysis covering release, separation, identification, and quantification to support clone selection, process development, and regulatory submission for glycoprotein therapeutics.
- 2-AB or procainamide labeling with HILIC-UPLC or CE-LIF profiling
- Site-specific glycan occupancy and heterogeneity by LC-MS/MS
- Biosimilar glycan comparability with statistical evaluation
- Glycan structure elucidation by exoglycosidase sequencing
Oxidation & Deamidation Analysis
Targeted detection and quantification of chemical degradation hotspots to support formulation development, stability protocol design, and acceptance criteria for product-related impurities.
- Site-specific oxidation and deamidation by LC-MS/MS peptide mapping
- Charge variant correlation by IEX and icIEF
- Forced degradation kinetics and Arrhenius modeling
- Specification justification based on degradation mechanism and clinical relevance
Beyond Glycosylation, Oxidation, and Deamidation
While glycosylation and common chemical degradations represent key PTM classes, protein therapeutics often exhibit a broader spectrum of modifications that can impact activity, stability, and immunogenicity. To address this complexity, our PTM characterization platform extends beyond dedicated modules to support the identification and analysis of additional modification types using flexible, method-driven workflows.
These include, but are not limited to, phosphorylation, acetylation, methylation, disulfide bond variants, C-terminal lysine processing, glycation, and other low-abundance or stress-induced modifications. Leveraging high-resolution LC-MS/MS, tailored enrichment or derivatization strategies, and advanced data interpretation pipelines, we enable both targeted and discovery-based PTM analysis at site-specific resolution.
Our integrated approach allows clients to comprehensively profile known and unknown PTMs, assess their structural and functional relevance, and incorporate these insights into process development, comparability studies, and regulatory documentation.
Analytical Methods Overview
Profacgen deploys a comprehensive suite of analytical techniques, selected and combined to interrogate your molecule's modification profile with maximum confidence and regulatory relevance.
- LC-MS/MS Peptide Mapping: High-resolution peptide mapping with multiple protease digestions localizes PTMs to specific amino acid residues and quantifies modification occupancy with high confidence, supporting both routine profiling and investigational root-cause analysis.
- Hydrophilic Interaction Liquid Chromatography (HILIC): HILIC separates released glycans by polarity, enabling quantitative glycosylation profiling, antennarity assessment, and sialylation level determination with high reproducibility for batch release and comparability.
- Ion-Exchange Chromatography (IEX) and icIEF: Charge-based methods detect and quantify deamidation, oxidation, and sialylation-induced charge variants, correlating macroscopic charge heterogeneity with specific chemical modifications to guide process understanding.
- Middle-Down and Top-Down Mass Spectrometry: Intact and subunit-level MS approaches preserve labile modifications and provide comprehensive PTM co-localization patterns inaccessible by bottom-up methods alone, offering a holistic view of modification landscapes.
Request a quote
Why Choose Us
- Comprehensive Multi-PTM Platform: We integrate glycan analysis, oxidation/deamidation profiling, disulfide mapping, and specialized modification screening into a unified program that eliminates analytical gaps and delivers a complete modification fingerprint.
- Regulatory-Aligned Methodologies: Our PTM assays are designed and executed to meet ICH Q6B, Q5C, FDA, and EMA expectations for product-related substance characterization, ensuring your data package supports IND, BLA, and biosimilar submissions.
- Site-Specific Localization and Quantification: We go beyond bulk detection to localize modifications to specific residues and quantify their occupancy, providing the mechanistic insight required for root-cause analysis, process optimization, and specification justification.
- Integrated Structural Biology Expertise: As part of our broader Structural & Physicochemical Characterization platform, PTM data seamlessly inform primary structure, higher-order structure, and impurity investigations—ensuring coherent interpretation across all quality attributes.
Representative Case Studies
Case 1: Biosimilar Glycosylation Comparability Assessment
Program Context:
A biosimilar development team required comprehensive glycosylation comparability data to demonstrate analytical similarity between their candidate monoclonal antibody and the reference product. Regulatory expectations mandated detailed glycan profiling, site-specific occupancy analysis, and quantitative comparison of critical glycan species such as afucosylated and high-mannose glycans.
Objective:
To generate a definitive glycosylation comparability package, including released glycan profiling, site-specific analysis, and relative abundance comparison, performed side-by-side under identical conditions with statistical evaluation.
Approach:
Profacgen performed N-glycan release with 2-AB labeling, followed by HILIC-UPLC profiling and CE-LIF confirmation for the biosimilar and reference product. Site-specific glycan occupancy was determined by LC-MS/MS peptide mapping of the conserved N-glycosylation site. Key glycan species were quantified as relative percentages and compared using predefined equivalence margins.
Outcome:
The biosimilar demonstrated a glycan profile statistically equivalent to the reference, with afucosylated and high-mannose species within biosimilarity margins. Site-specific occupancy was comparable, and no aberrant glycan structures were detected. The glycosylation package supported successful regulatory submission and progression to clinical comparability studies.
Case 2: Root-Cause Investigation of Charge Variant Increase During Stability
Program Context:
A pharmaceutical company observed an unexpected increase in acidic charge variants during long-term stability studies of a therapeutic antibody, threatening shelf-life claims and release specifications. The team required identification of the specific chemical degradation mechanism to guide reformulation or process adjustment.
Objective:
To identify and localize the PTM(s) responsible for the acidic charge variant increase, quantify their rate of formation, and recommend formulation or process modifications to mitigate the degradation.
Approach:
We subjected stressed and control samples to icIEF for charge variant quantification, then performed LC-MS/MS peptide mapping with multiple protease digestions to localize modifications. IEX fractionation isolated acidic species for targeted analysis, and forced degradation studies at elevated pH and temperature accelerated the reaction for mechanistic insight.
Outcome:
Peptide mapping identified a specific asparagine residue undergoing deamidation at an accelerated rate in the original formulation buffer. Isomerization to the isoaspartate form was confirmed, explaining the charge shift. A reformulated buffer at optimal pH reduced the deamidation rate by 60% during accelerated testing, supporting an extended shelf-life claim and revised release specifications.
Get a Project Assessment
Frequently Asked Questions (FAQs)
Q: What is PTM characterization?
A: Post-translational modification (PTM) characterization is the comprehensive analytical process of identifying, localizing, and quantifying chemical alterations that occur to proteins after ribosomal translation. These modifications—including glycosylation, oxidation, deamidation, phosphorylation, acetylation, and disulfide bond formation—are not genetically encoded and can profoundly influence therapeutic efficacy, stability, half-life, and safety. Rigorous PTM characterization establishes critical quality attributes (CQAs), ensures batch-to-batch consistency, and provides the regulatory evidence required for IND, BLA, and biosimilar submissions.
Q: Why is glycosylation profiling important?
A: Glycosylation is one of the most structurally complex and functionally consequential post-translational modifications in biologics. Glycan composition directly modulates antibody effector functions such as antibody-dependent cellular cytotoxicity (ADCC) and complement-dependent cytotoxicity (CDC), influences serum half-life through FcRn interactions, and affects immunogenic potential. Because glycosylation patterns vary with cell line, culture conditions, and purification process, comprehensive profiling is essential for clone selection, process control, and demonstrating comparability. Regulatory agencies classify glycosylation as a critical quality attribute and require detailed characterization to ensure therapeutic performance and consistency.
Q: What techniques detect oxidation and deamidation?
A: Oxidation and deamidation are detected through orthogonal analytical techniques that combine separation, mass spectrometry, and charge analysis. High-resolution LC-MS/MS peptide mapping localizes these modifications to specific amino acid residues by detecting characteristic mass shifts—such as +0.984 Da for deamidation and +16 Da for methionine oxidation—and quantifying their relative abundance. Ion-exchange chromatography (IEX) and imaged capillary isoelectric focusing (icIEF) detect the charge alterations associated with deamidation and oxidation, providing complementary bulk quantification. Hydrophobic interaction chromatography (HIC) can resolve oxidized species by altered surface hydrophobicity, while forced degradation studies validate method sensitivity and establish kinetic profiles.
Q: Can PTMs affect immunogenicity?
A: Yes, post-translational modifications can significantly influence the immunogenicity of protein therapeutics. Certain modifications—such as aggregation-prone oxidation, non-human glycan structures, or deamidation-induced conformational changes—can expose cryptic epitopes or create neo-epitopes that are recognized as foreign by the immune system. Additionally, modifications that alter protein folding or stability may increase the propensity for aggregate formation, which is a well-established risk factor for anti-drug antibody (ADA) development. Comprehensive PTM profiling identifies these liability hotspots early, enabling informed clone selection, process optimization, and formulation design to mitigate immunogenicity risks before clinical testing.
Q: Is PTM characterization required for biosimilars?
A: Yes, PTM characterization is a mandatory component of biosimilar development programs. Regulatory agencies—including the FDA and EMA—require detailed comparability data demonstrating that the biosimilar candidate possesses post-translational modification profiles highly similar to the reference product. Glycosylation patterns, oxidation levels, deamidation rates, and other modifications must be quantified and compared using orthogonal methods with predefined equivalence margins. Because PTMs directly impact efficacy, safety, and immunogenicity, close alignment of modification profiles provides essential evidence for the analytical similarity package and supports the totality-of-evidence approach required for biosimilar approval.
Q: How long does comprehensive PTM analysis take?
A: A comprehensive PTM characterization program typically requires 4–6 weeks from sample receipt to final report, though timelines vary based on the complexity of the analytical scope and the number of modifications targeted. Standard glycosylation profiling by HILIC-UPLC or CE-LIF can be completed within 2–3 weeks, while site-specific peptide mapping with LC-MS/MS for oxidation, deamidation, and disulfide bond analysis may extend to 3–4 weeks. Programs incorporating multiple orthogonal techniques, forced degradation kinetics, or extensive biosimilar comparability studies may require 6–8 weeks. We provide detailed project timelines during initial consultation and offer expedited services for time-critical regulatory submissions.
Reference:
- Bagwan N, El Ali HH, Lundby A. Proteome-wide profiling and mapping of post translational modifications in human hearts. Sci Rep. 2021;11(1):2184. doi:10.1038/s41598-021-81986-y

 Figure 1. An example of PTM identifications in human hearts. (Adapted from Bagwan et al., 2021)
Figure 1. An example of PTM identifications in human hearts. (Adapted from Bagwan et al., 2021)